Extra: AlphaFold-Multimer
On October 4th, 2021, DeepMind released the AlphaFold-Multimer publication on bioRxiv, closely followed by updated code and models on GitHub. AF-Multimer is an adaptation of the original architecture, but now specifically trained to more accurately model protein-protein interactions. This page gives a summary of the implications for AlphaFold on the HPC when you want to run these specialized models.
1. Change the FASTA file
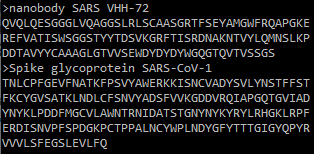
The proteins that you want to fold with AlphaFold-Multimer should be placed in a single FASTA file, with one entry per protein.
2. Adaptations to the job script
The main change to the job script is to set the used models to multimer instead of monomer_ptm. Other than that, you can specify that input proteins are all from prokaryotes. If not, do not include this line.

3. GPU limits + running time
The maximum sequence length that the GPU can fit remains the same, taking the sum of the sequence length for all proteins as a total. Running time will be higher, as a separate MSA search is done for each protein in input.
4. Changes in the output
The output remains largely the same. You will now have an MSA per protein specified. Also, the .pdb files contain multiple chains.
5. PTM vs pLDDT
Note that the ranking of different predicted structures is no longer done based on pLDDT for AlphaFold-Multimer, but on the PTM, as we are mainly interested in how accurately the interactions are modeled. The PTM for each model can be found in the ranking_debug.json output file.